Upstream processing
Common problems
Batch-to-batch variability is often measured extensively but rarely explained
Scientists and engineers involved in upstream processing typically face several difficulties:

Our solutions
Software license - Bioreactor software
Our software was developed in collaboration with Sanofi during the CALIPSO project. It features a user-friendly interface to ensure data completeness and re-usability, perform calculations (mu, Qp, Qs, …) in a few clicks, compare data and generate automatic reports. This allows significant time saving and reduces the risk of errors, thus allowing scientists and engineers to focus on their core activities.
Our software also provides mechanistic simulation capabilities allowing to gain process understanding and reduce the number of experiments.
Comprehensive support - Automated data structuration
Our software already contains some data import functionalities. But we know each team works in a unique environment, with specific needs and constraints. To bridge the gap between flexibility and standardization, we offer the development of custom scripts to automate the import of experimental data in our structured format. The scripting tool is in an open format such that data scientists can also develop their own scripts as needed.
Because data standardization is a pillar of exploiting your data, may it be through Ypso-Facto software or your own solutions, automated data structuration opens a wide range of possibilities.
Collaborative partnership - Extended team
Our team of experts provides comprehensive upstream process development services. We work alongside your scientists and engineers, challenging assumptions and complementing internal initiatives.
For instance, our “EXPLORE” package is a good way to kick-start a new journey towards the implementation of simulation tools. We demonstrate the benefits of modeling approaches on a concrete example and provide a methodology to be re-used on future projects.
Case studies
Process: CHO cell culture
Product: mAb
Results: This work aimed at comparing our mechanistic model with models traditionally used in the literature. We fitted on batch data
- our mechanistic model (blue dashed line),
- a Monod model with one route only considering μ1=f(Glc, Gln, NH4+, Lac) (red dotted line),
- and a Monod model with three routes considering μ1=f(Glc, Gln, NH4+, Lac), μ2=f(Glc, Lac), μ3=f(Lac), (green dash-dot line).
Then, we compared predictions of the 3 models on fed-batch data in terms of VCD, glucose, lactate, glutamine, ammonia and IgG concentrations (Fig. 1). It was found that our model outperforms the Monod models. Indeed, with the Monod model considering one route only, the VCD is significantly underestimated since glutamine is completely depleted. With the Monod model considering three routes, the VCD continues to increase even at the end of the culture since glucose is still present. No Monod model could be found to represent this set of data.
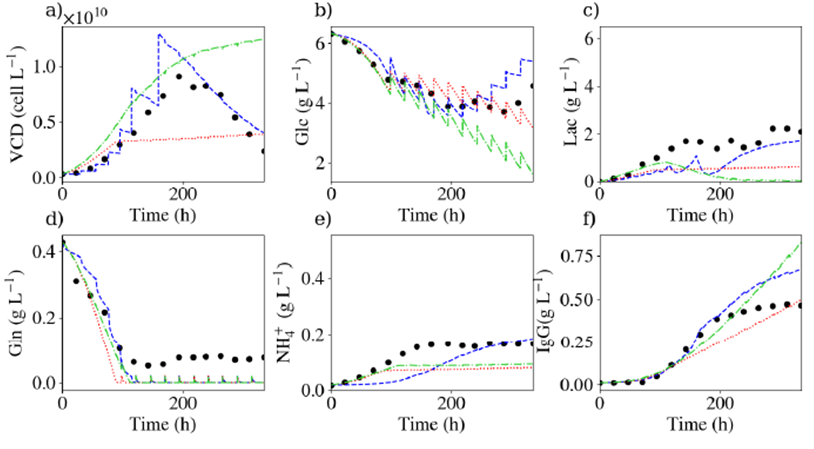
Fig1: comparison between experimental (dots) and simulation (lines) results for commonly measured quantities
These results suggest the need for a more detailed description of the physico-phenomena at state than the classical Monod models. In particular, our model allows simulating the concentrations of all amino acids. This allowed to identify that some amino acids like Asparagine (Asn) were limiting (Fig. 2). Performing fed-batch experiments with the addition of well targeted amino acids allowed to significantly improve process performances.
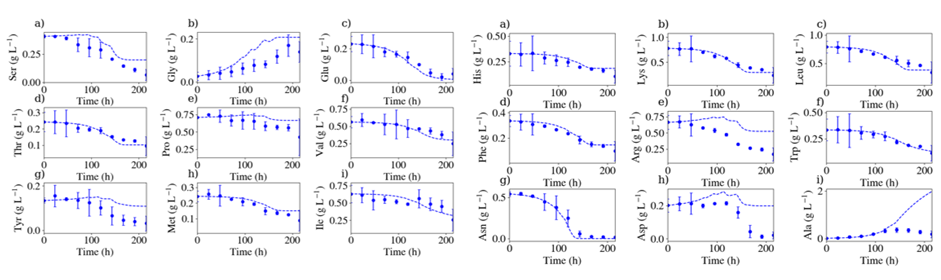
Fig2: comparison between experimental (dots) and simulation (lines) results for amino acids
Key message: our mechanistic model for mAb production does not only account for glucose, glutamine, lactate and ammonia but also all amino acids.
Further reading: mAb production kinetics in CHO batch culture: exploring extracellular and intracellular dynamics
Process: E Coli and CHO
Product: antibodies, vaccines, GFP
Results: This work aimed at automatically structuring experimental data from various sources. The workflow was as follows
- Successful connection to a datalake shared by 2 sites of Sanofi (364 bioreactor experiments on 8 different projects) and possibility to apply search criteria to find the experiments of interest (Fig. 1)
- Automatic import of online, atline, offline and set points data of selected experiments in the bioreactor module
- For data not stored in the datalake, development of custom scripts to achieve automatic import in the bioreactor module
- After import, access to standard functionalities of the bioreactor module including calculation of quantities of interest like mu, Qs, Qp, easy comparison of experiments, automatic Word report of experiments
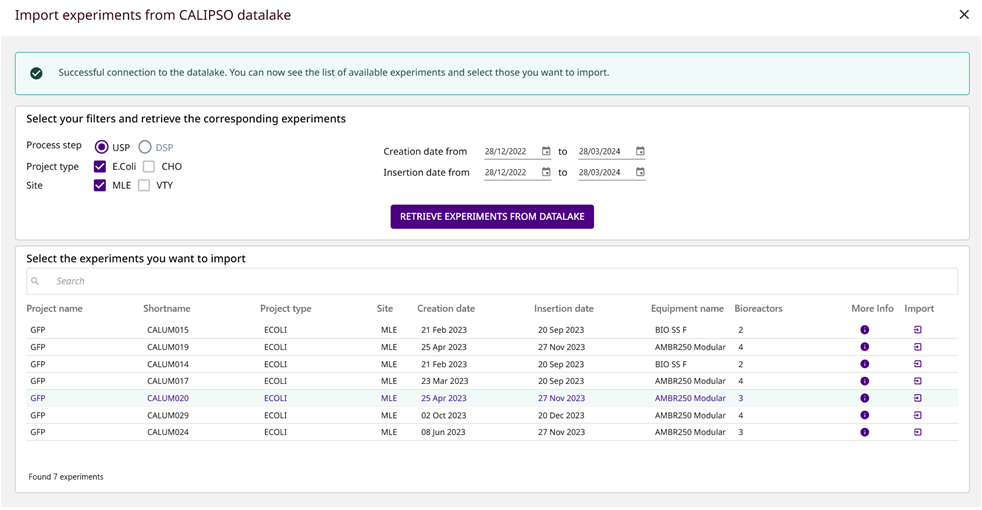
Fig1: Connection to the datalake to retrieve experiments
Key message: it is straightforward to automate data structuration and standardization by developing connectors between existing excel files and the bioreactor module
Process: E Coli
Product: GFP
Results: An experimental plan investigating the impact of temperature, pH and feeding coefficient was performed. Five experiments were used for fitting purposes. The following results were simulated: product, biomass, glucose, glutamate, formate, ammonia, acetate, CO2 and found in very good agreement with experimental data (Fig. 1).
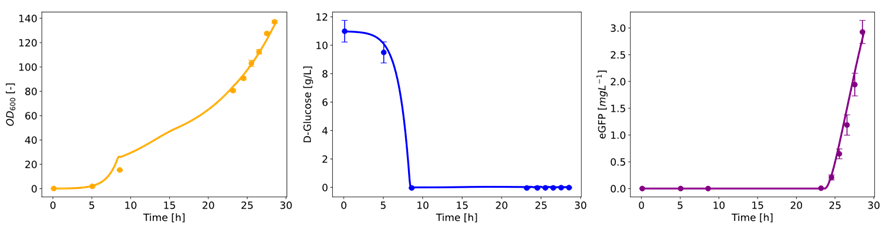
Fig. 1: comparison between experimental (dots) and simulation (lines) results for the optical density, glucose concentration and GFP concentration of the reference experiment.
Then, the model was used to predict 7 additional experiments in order to verify model accuracy. It was found that the model is capable of describing the entire set of data (Fig. 2).
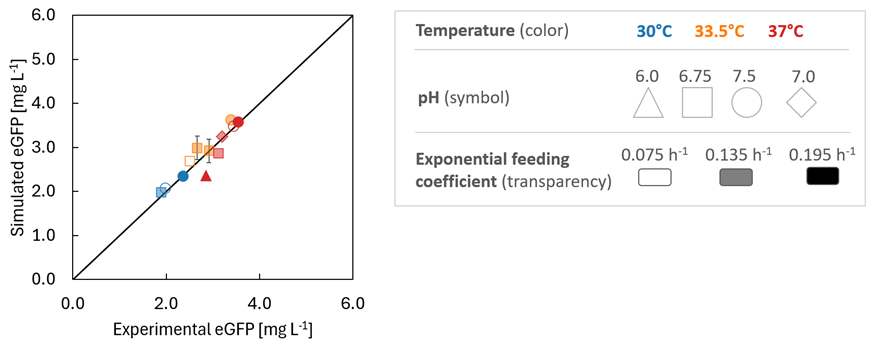
Fig. 2: Comparison between experimental and simulated results for the entire data set
Finally, the model was used to predict the impact of selected process parameters: temperature, pH, feeding rate (Fig. 3).
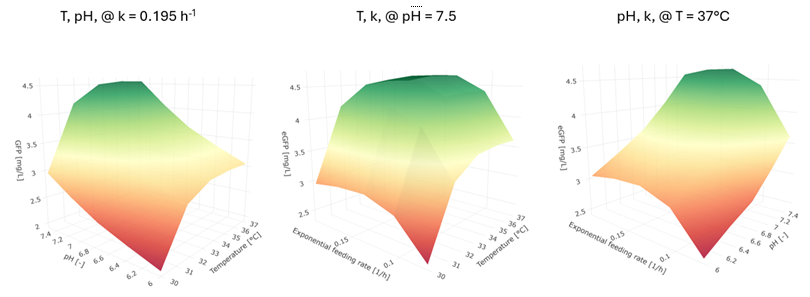
Fig. 3: Simulated impact of the temperature, pH and feeding rate
Key message: our mechanistic model can be used to predict the behavior of E. Coli cultures
Your questions, answered
The data structuration part of the software can be used for any type of microorganism. The mechanistic model is available for both CHO and E Coli cultures.
The number of experiments depends on the objectives and constraints of the project. To give a rough idea, with a dozen of bioreactors, a first version of the model can be obtained.
The mechanistic model embedded in the bioreactor software can predict the impact of a number of operating parameters including:
- the concentrations of glucose, glutamine and all amino acids in the culture medium
- the concentrations of glucose, glutamine and all amino acids in the feed
- the feeding strategy (moment of addition, volume of addition, flowrate, etc)
- a switch from batch to perfusion culture
Want to know more?
Don't hesitate to contact us!
